- Home
- Labs
- Pierce Lab
- Projects
- High-throughput Functional Studies of Sequence Variants
Obtaining a genetic diagnosis for inherited retinal diseases is increasingly important, but bioinformatic predictions of pathogenicity for DNA variants of unknown significance (VUS) are imperfect– a major bottleneck in interpreting human genetic variation.
Purpose: Develop higher-throughput cell-biological methods to evaluate variants for pathogenicity with higher confidence – a “functional genomics” approach.
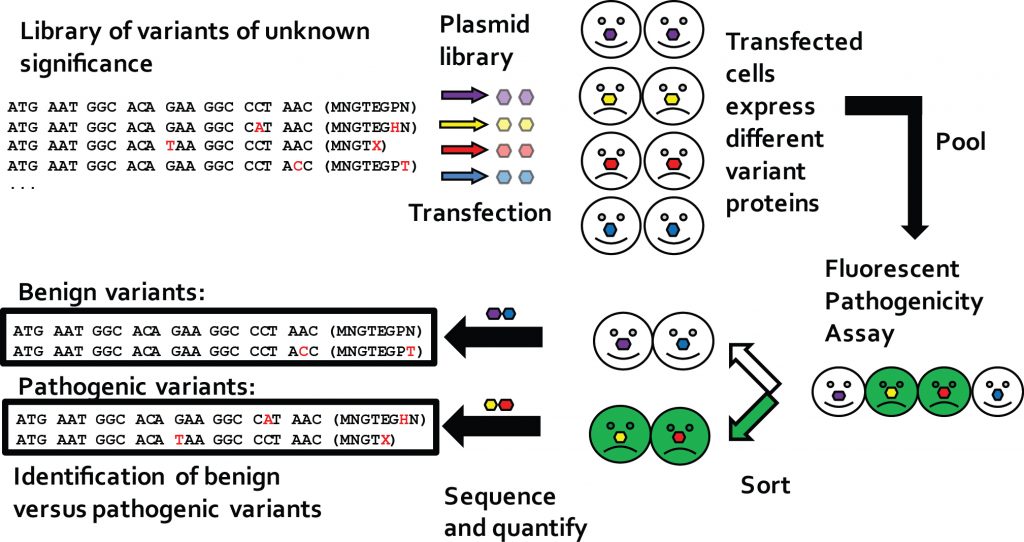
A library approach was developed to efficiently detect pathogenic rhodopsin (RHO) variants that fail to express on the cell surface.
This approach, while initially focused on RHO, is meant to demonstrate a streamlined, general method for detecting pathogenic VUS.