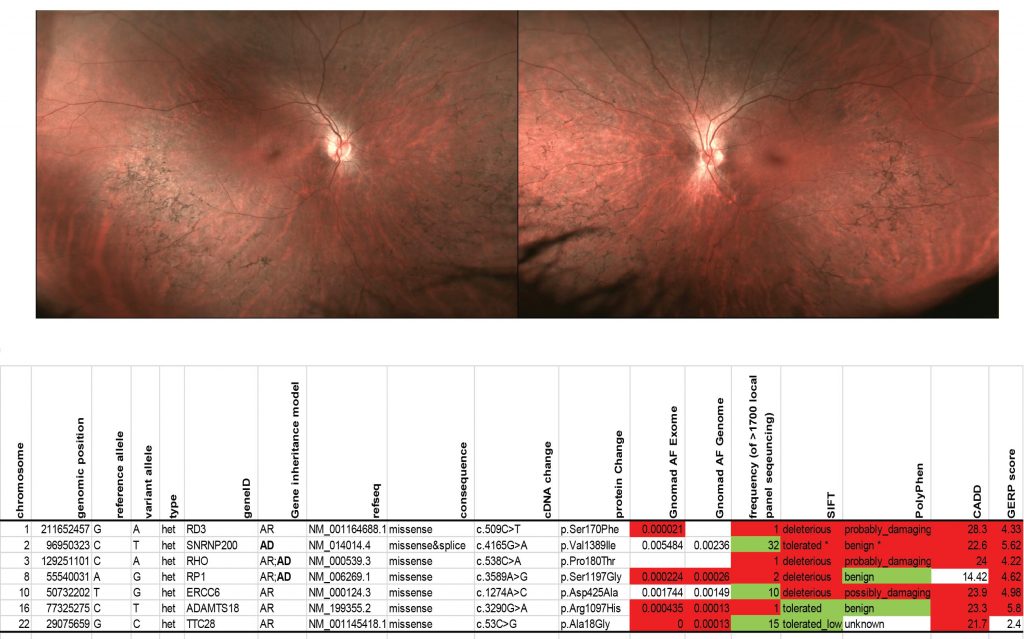
Our work focuses on using techniques of functional genomics to better understand which DNA variants found in patients with inherited retinal disorders are pathogenic. In fact, characterizing the pathogenicity of DNA sequence variants of unknown significance (VUS) is a major bottleneck in human genetics, and is increasingly important in determining which patients with inherited retinal diseases could benefit from gene therapy. For example, a library of 210 rhodopsin (RHO) variants from literature and in-house genetic diagnostic testing was created to efficiently detect pathogenic RHO variants that fail to express on the cell surface. The resulting dataset characterized surface expression of every RHO library variant with a high degree of reproducibility, recategorizing 37 variants. Three retinitis pigmentosa pedigrees were solved by identifying VUS which showed low expression levels. Results were validated across multiple assays and correlated with clinical disease severity. This is just one example of how functional genomics can be applied to understanding VUS in IRDs.
Newer projects include base editing to correct missense mutations which cause inherited retinal diseases. We also work on collaborative projects to improve the delivery of gene therapy vectors to the retina in higher animals.
